The coronavirus in a tiny drop
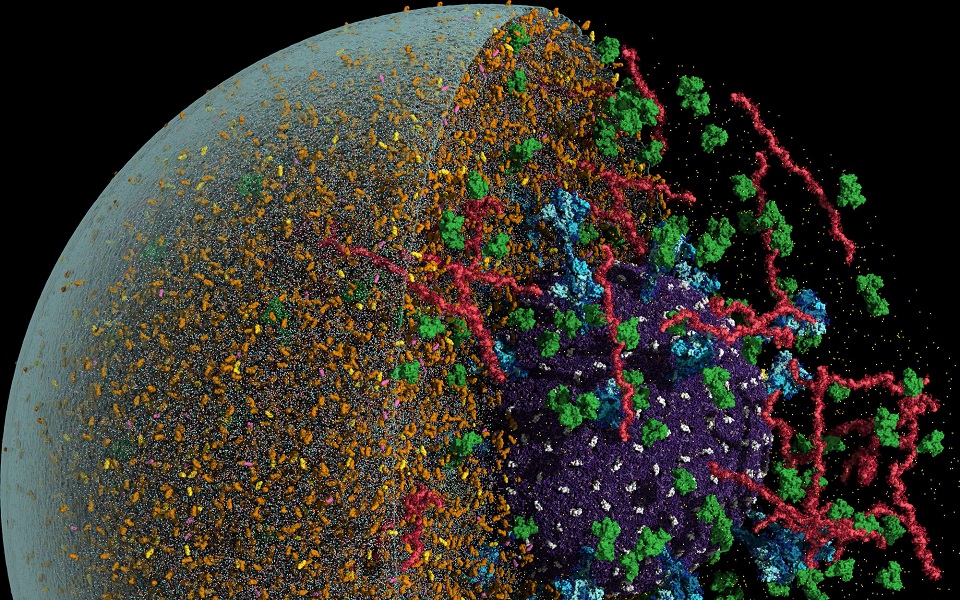
To better understand the coronavirus’s journey from one person to another, a team of 50 scientists has for the first time created an atomic simulation of the coronavirus nestled in a tiny airborne drop of water.
To create the model, the researchers needed one of the world’s biggest supercomputers to assemble 1.3 billion atoms and track all their movements down to less than a millionth of a second. This computational tour de force is offering an unprecedented glimpse at how the virus survives in the open air as it spreads to a new host.
“Putting a virus in a drop of water has never been done before,” said Rommie Amaro, a biologist at the University of California San Diego who led the effort, which was unveiled at the International Conference for High Performance Computing, Networking, Storage and Analysis last month. “People have literally never seen what this looks like.”
How the coronavirus spreads through the air became the subject of fierce debate early in the pandemic. Many scientists championed the traditional view that most of the virus’s transmission was made possible by larger drops, often produced in coughs and sneezes, which usually travel only a few feet before dropping to the floor.
But epidemiological studies showed that people with COVID-19 could infect others at a much greater distance: Just talking in a poorly ventilated space like a bar, church or classroom was enough to spread the virus.
Those findings pointed to much smaller drops, called aerosols, as important vehicles of infection. Scientists define droplets as having a diameter greater than 100 micrometers, or about four-thousandths of an inch. Aerosols are smaller — in some cases so small that only a single virus can fit inside them. And thanks to their minuscule size, aerosols can drift in the air for hours.
Viruses cannot survive forever in aerosols, though. Researchers often find that viruses collected from the air have become so damaged that they can’t infect cells anymore. It’s possible that as the aerosols evaporate, the air destroys the virus’s molecular structure. Or the chemistry inside the tiny drop may become too hostile for them to survive.
“At this point, we don’t understand how that happens,” said Linsey Marr, a professor of civil and environmental engineering at Virginia Tech who was not involved in the new study. Microscopes that can capture detailed images of what goes on inside a virus-laden aerosol have yet to be invented.
In March 2020, Amaro and her colleagues decided the best way to open this black box was to build a virus-laden aerosol of their own.
The researchers started by creating a model of the coronavirus, known as SARS-CoV-2, from 300 million virtual atoms. They combined thousands of fatty acid molecules into a membrane shell, then lodged hundreds of proteins inside. Some of these proteins are important because they keep the membrane intact. Others, called spike proteins, form flowerlike structures that rise far above the surface of the virus. The tips of the spikes sometimes spontaneously flick open, allowing the virus to latch on to a host cell and invade.
After building their virus, Amaro and her colleagues made an aerosol to put it in. Using 1 billion atoms, they created a virtual aerosol measuring a quarter of a micrometer in diameter, less than a hundredth the width of a strand of human hair.
The researchers could not simulate the aerosol as a blob of pure water, however. When an aerosol breaks free from the fluid in our lungs, it brings along a stew of other molecules from our bodies.
That stew includes mucins: long, sugar-studded proteins from a lung’s lining. Aerosols also carry molecules called surfactants, which keep the delicate branches of our airways from sticking together. Charged atoms, like calcium and magnesium, fly around the aerosol, exerting powerful forces on molecules they encounter.
Once the virus was loaded into an aerosol, the scientists faced the biggest challenge of the project: bringing the drop to life. Amaro and her colleagues calculated the forces at work across the entire aerosol, taking into account the collisions between atoms as well as the electric field created by their charges. They determined where each atom would be four-millionths of a billionth of a second later.
To carry out this vast set of calculations, the researchers had to take over the Summit Supercomputer at the Oak Ridge National Laboratory in Tennessee, the second-most-powerful supercomputer in the world. Because the machine was in high demand, they could run their simulation only a few times. “We only have so many shots to actually see if we can get this thing to actually fly,” Amaro said.
The first run was a disaster. Tiny flaws in their model caused the virtual atoms to crash into one another, and the aerosol instantly blew apart. “It basically explodes,” Amaro said.
After half a dozen rounds of adjustments, the aerosol became stable. The researchers ran the calculations all over again to see what happened inside the aerosol an instant later. All told, they created millions of frames of a movie capturing the aerosol’s activity for ten-billionths of a second.
“While molecular modeling is not a new thing, the scale of this is next level,” said Brian O’Flynn, a postdoctoral research fellow at St. Jude Children’s Research Hospital who was not involved in the study.
The buzzing activity Amaro and her colleagues witnessed offered clues about how viruses survive inside aerosols. The mucins, for example, did not just wander idly around the aerosol. The negatively charged mucins were attracted to the positively charged spike proteins.
Amaro speculated that the mucins act as a shield. If the virus moves too close to the surface of the aerosol, the mucins push them back in, so that they aren’t exposed to the deadly air. “What we think is that it’s actually covering itself in these mucins, and that’s acting like a protective coating for it during flight,” Amaro said.
This discovery may help explain how the delta variant became so widespread. Delta’s spike proteins have a more positive charge than those on earlier forms of the coronavirus. As a result, mucins huddle more closely around them. That attraction could potentially make the mucins a better shield.
Every now and then, one of the simulated coronaviruses flipped open a spike protein, surprising the scientists. “The delta variant opens much more easily than the original strain that we had simulated,” Amaro said.
Once a coronavirus enters someone’s nose or lungs, the delta spike’s wide opening may make it better at infecting a cell. But Amaro suspects that it’s bad for a coronavirus to open a spike protein when it’s still inside an aerosol, perhaps hours away from infecting a new host. “If it opens too soon, it could just fall apart,” Amaro said.
Some of the molecules that are abundant inside aerosols may be able to lock the spike shut for the journey, she said. Certain lung surfactants can fit into a pocket on the surface of the spike protein, preventing it from swinging open.
To test that idea and explore others, Amaro and her colleagues are stretching out the time frame of their simulation 100 times, from ten-billionths of a second to a millionth of a second. They’ll also investigate how the acidity inside an aerosol and the humidity of the air around it may change the virus.
Amaro and her colleagues are making plans to build an omicron variant next and observe how it behaves in an aerosol. They want to wait for structural biologists to work out the three-dimensional shape of its spike proteins before getting started. But just looking at the early findings about omicron, Amaro already sees one important feature: “It is even more positively charged,” she said.
Because omicron’s spike proteins are even more positively charged than delta’s, it may build a better mucin shield in aerosols. And that may help make it even more transmissible.
Marr said the simulation might eventually allow scientists to predict the threat of future pandemics. They could build atomic models of newly discovered viruses and put them into aerosols to watch them behave.
“This has implications for understanding emerging viruses that we don’t yet know about,” Marr said. “There’s still a long way to go to get there,” she said, “but this is definitely a big first step.” [Science Times]
This article originally appeared in The New York Times.